Elevate your single-cell research with CosMx™ SMI
A Spatial Multiomics Single-Cell Imaging Platform
CosMx SMI is the first high-plex in situ analysis platform to provide spatial multiomics with formalin-fixed paraffin-embedded (FFPE) and fresh frozen (FF) tissue samples at cellular and subcellular resolution. CosMx SMI enables rapid quantification and visualization of up to 6,000 RNA and 64 validated protein analytes. It is the flexible, spatial single-cell imaging platform that will drive deeper insights for cell atlasing, tissue phenotyping, cell-cell interactions, cellular processes, and biomarker discovery.
The CosMx™ SMI and decoder probes are not offered and/or delivered to the Federal Republic of Germany for use in the Federal Republic of Germany for the detection of cellular RNA, messenger RNA, microRNA, ribosomal RNA and any combinations thereof in a method used in fluorescence in situ hybridization for detecting a plurality of analytes in a sample without the consent of the President and Fellows of Harvard College (Harvard Corporation) as owner of the German part of EP 2 794 928 B1. The use for the detection of cellular RNA, messenger RNA, microRNA, ribosomal RNA and any combinations thereof is prohibited without the consent of the President and Fellows of Harvard College (Harvard Corporation).
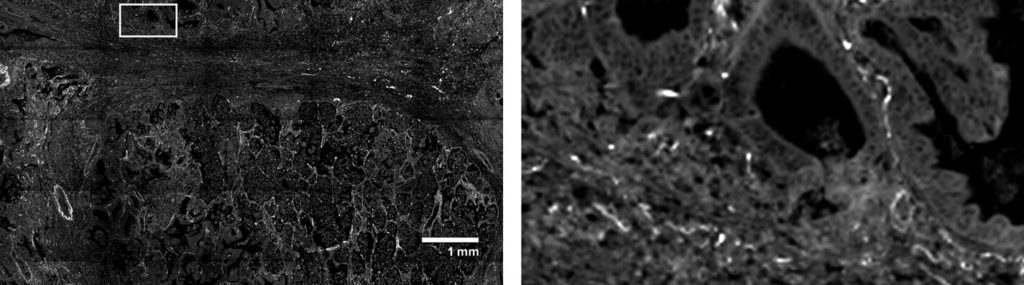



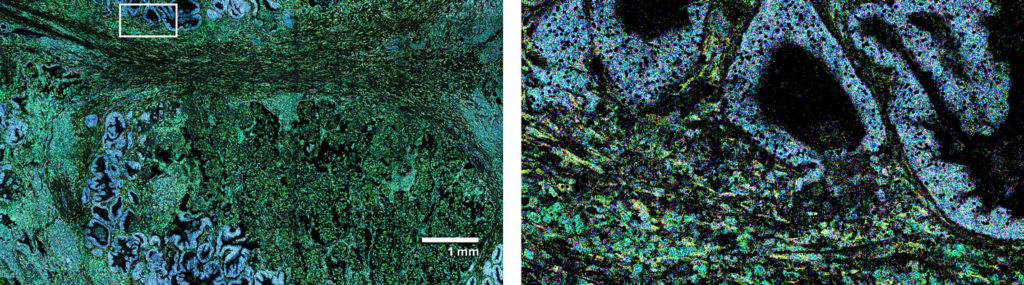

“We are impressed with ease of use with which the CosMx SMI enabled single cell spatial transcriptomics across various tissue types, even in archival FFPE tissue. The high-plex protein assay makes the CosMx SMI a true multiomic solution. The CosMx SMI’s large capture area allows analysis from valuable clinical trial samples using tissue microarrays, enabling characterization of 100s of samples weekly.”

Dr. Nigel B Jamieson
Professor of Surgery, University of Glasgow, UK
“At last count, we have processed 160 samples with the CosMx SMI, including tumor and mouse brain samples. This technology uncovers vital insights into cell biology, enhancing our understanding of both malignant and normal tissues. It is crucial for creating spatial biomarkers and improving patient outcomes. Plus, the equipment and protocols are reliable and user-friendly.”

Dr. Aubrey Thompson
Mayo Clinic Comprehensive Cancer Center, Florida
“The CosMx SMI complemented our efforts to generate organ atlases of the healthy human system together with our partners of the Human Cell Atlas project. It allowed us to visualize single-cell reference maps directly in organs. The scale of 1000-plex gene panel with image-guided cell segmentation makes the CosMx SMI, a unique platform for spatial genomics!”

Dr. Holger Heyn
Team Leader, Single Cell Genomics Group, CNAG, Spain
“The CosMx SMI instrument has become integral to our laboratory’s future direction, and the responsiveness and skill of the customer support for it and the AtoMx SIP has been absolutely fantastic.”

Dr. Grant Kolar
Director, Research Microscopy and Histology Core, St. Louis University School of Medicine
Be the first to get…
Comprehensive answers with 6000-plex RNA assays in a single experiment
Get single-cell gene expression in challenging FFPE tissues with spatial context.
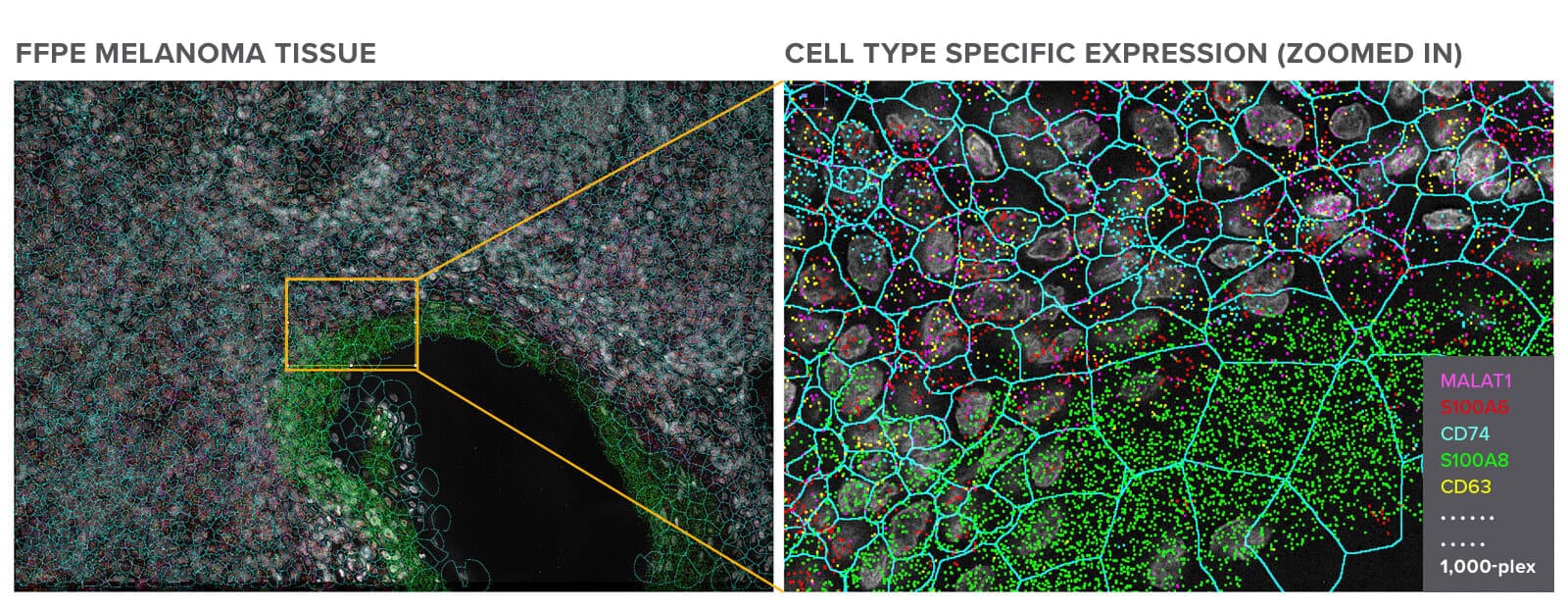
Melanoma FFPE tissue probed with 1000-plex RNA panel to detect spatial localization of transcripts in intact tissue.
Detect low expressing genes with highly sensitive CosMx SMI
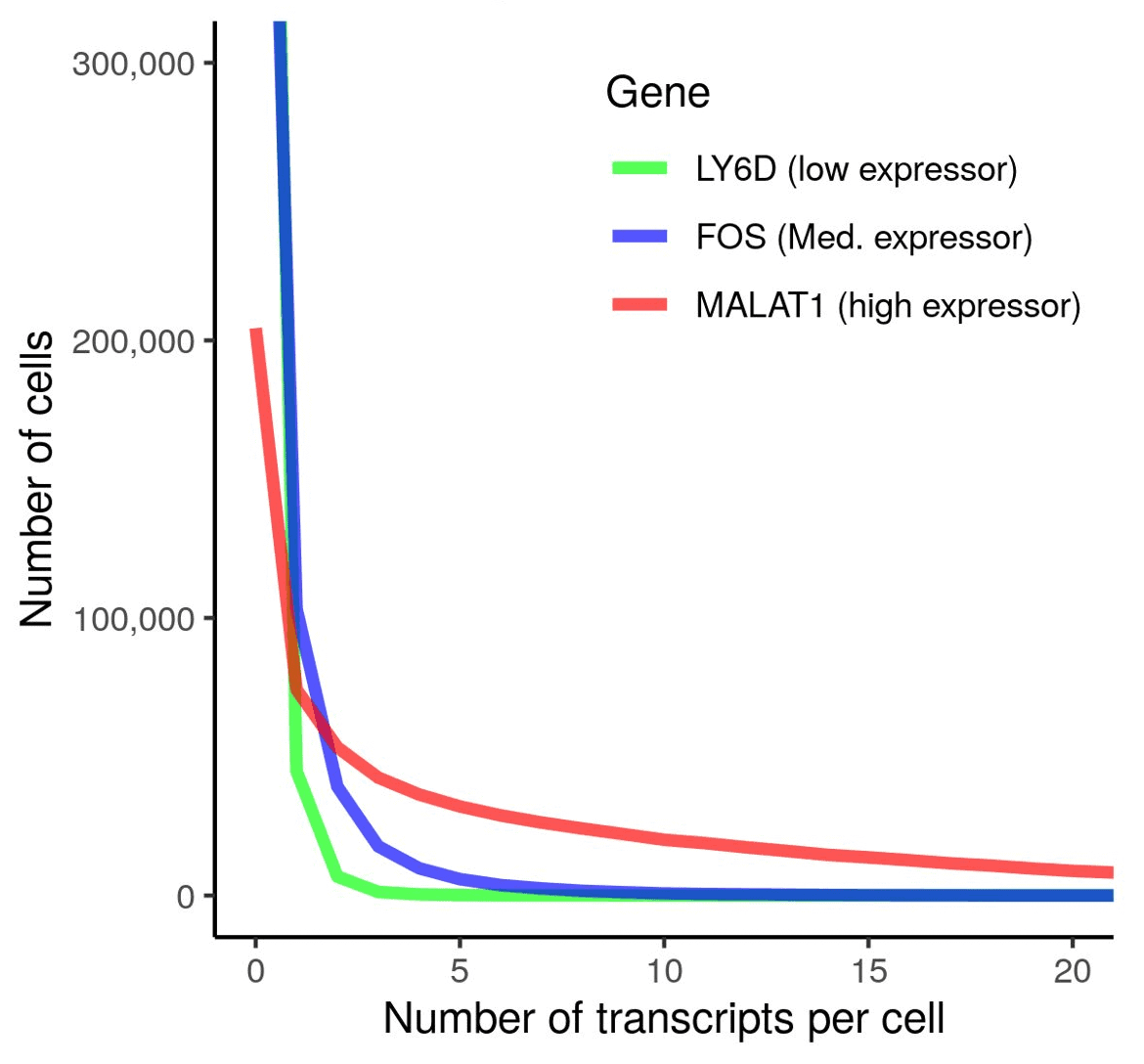
| Genes | Expression level | Copies per cell (Max.) |
|---|---|---|
| MALAT1 | High | 256 |
| FOS | Medium | 40 |
| LY6D | Low | 24 |
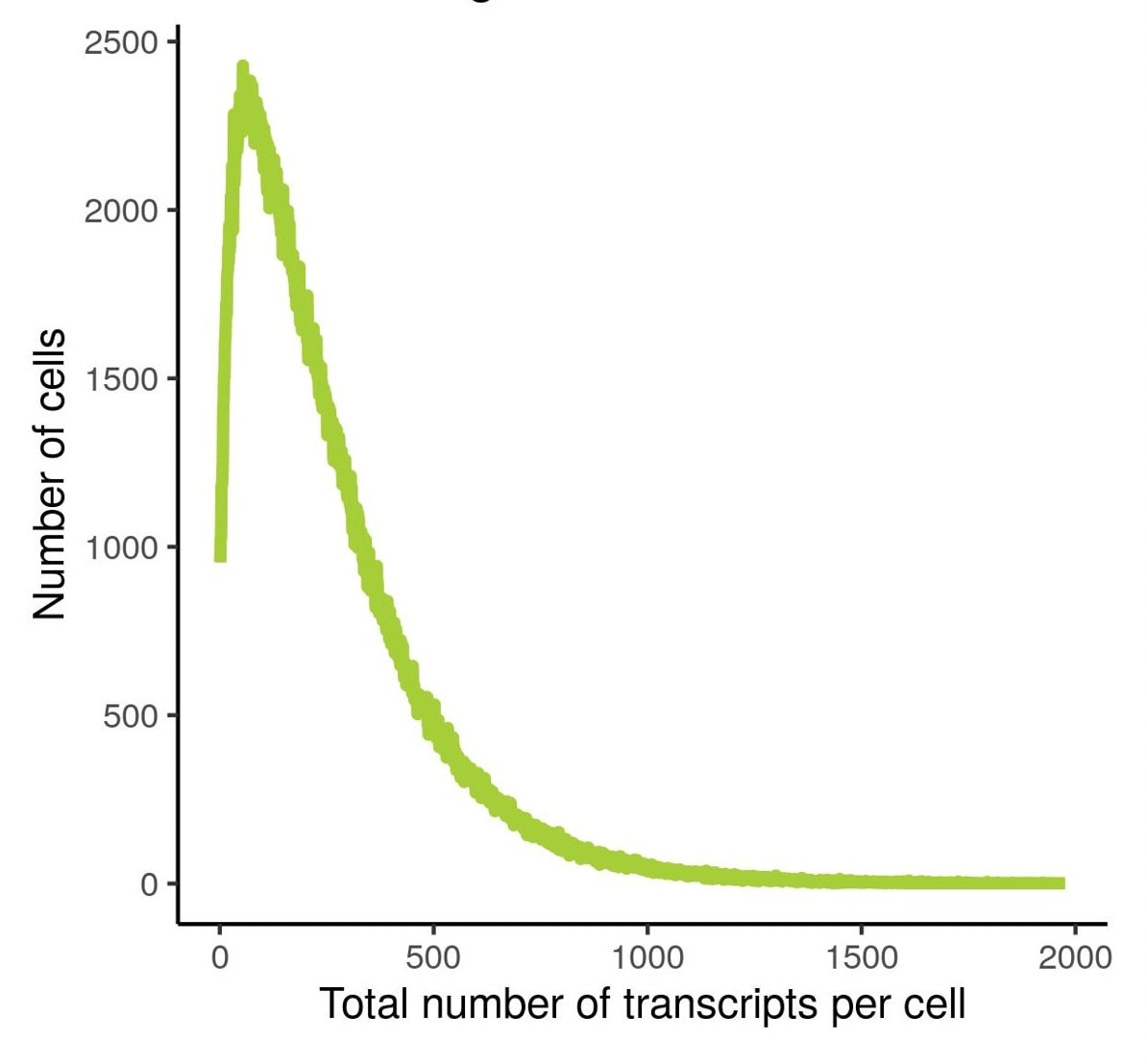
Total number of cells analyzed = 800,327
Total transcripts detected = 259,604,214
Number of transcripts / cell (Max) = 2000
Number of transcripts / cell (Mean) = 257
Shown here are the transcripts detected per cell in non-small cell lung cancer (NSCLC) FFPE tissue probed with a 1000-plex gene panel (left) and total transcript distribution in the NSCLC tissue (right). CosMx SMI offers high sensitivity and high dynamic range to capture low copy number gene transcripts at the single cell level.
Multi-modal approach for true single-cell segmentation
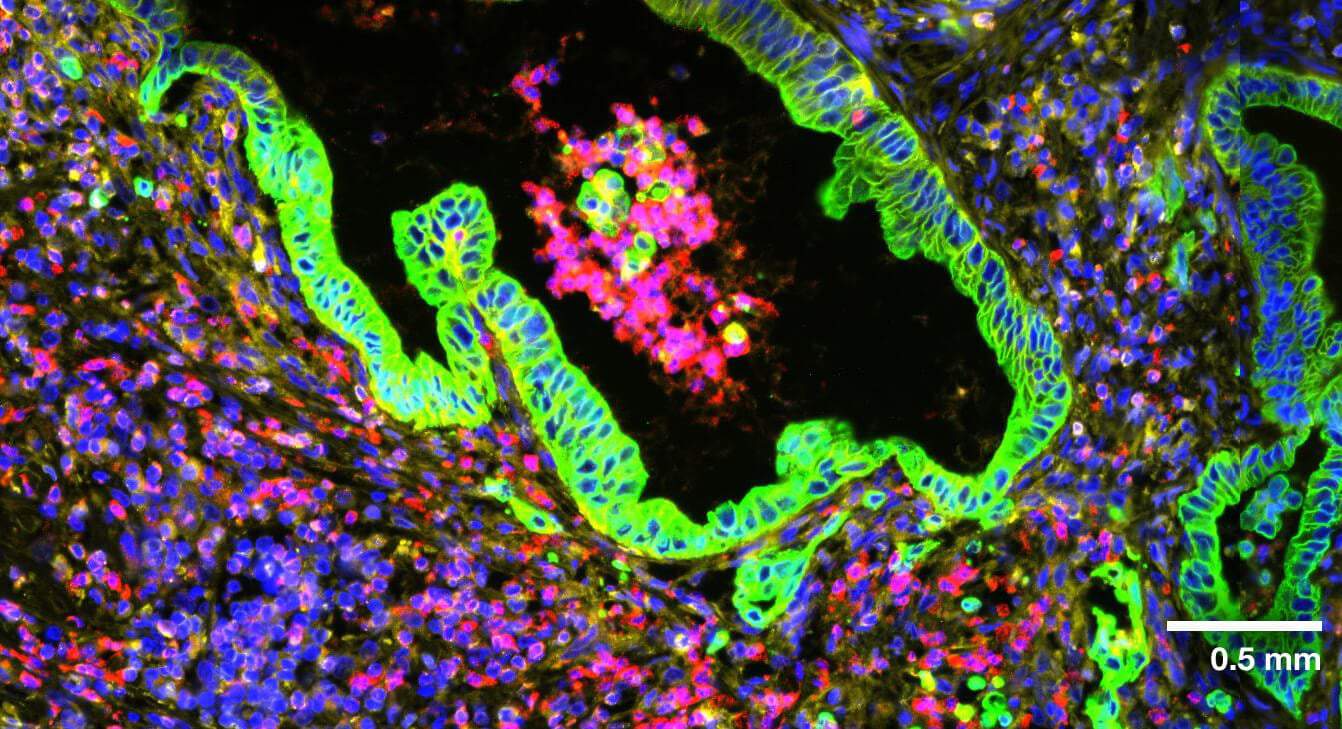

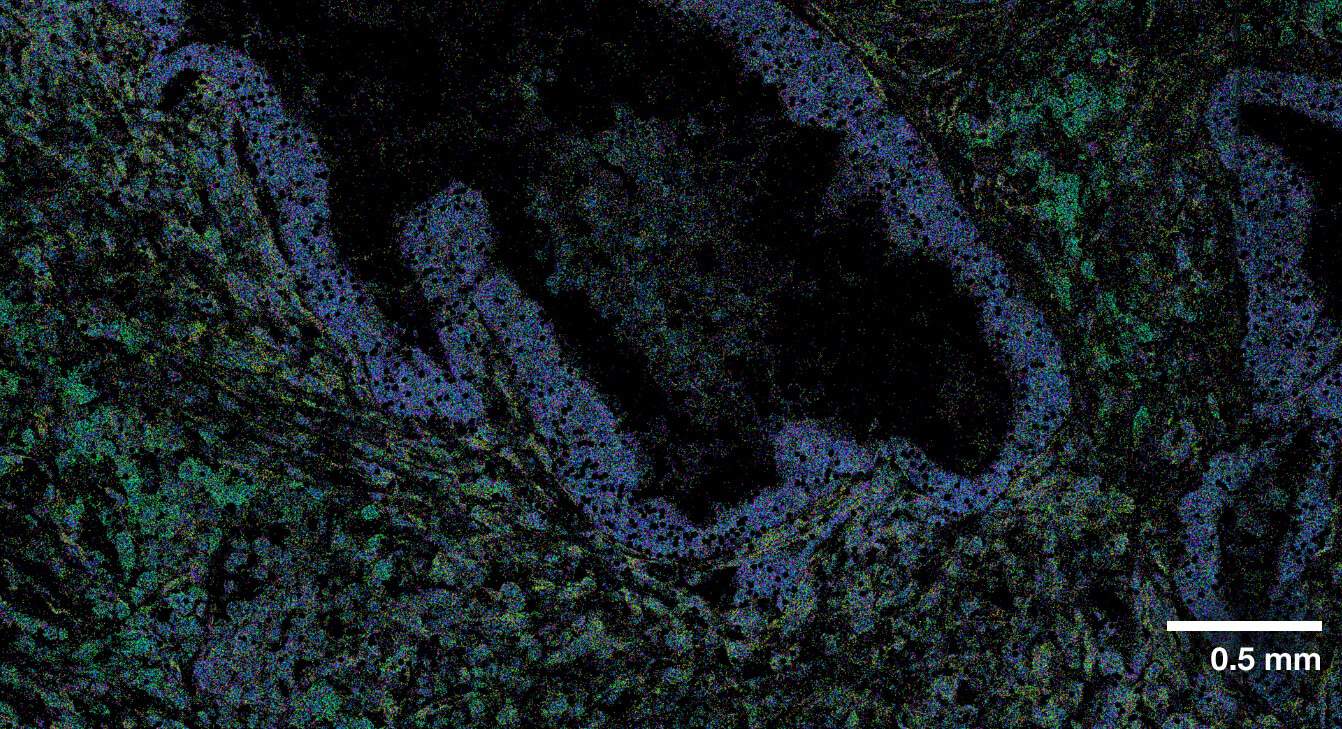
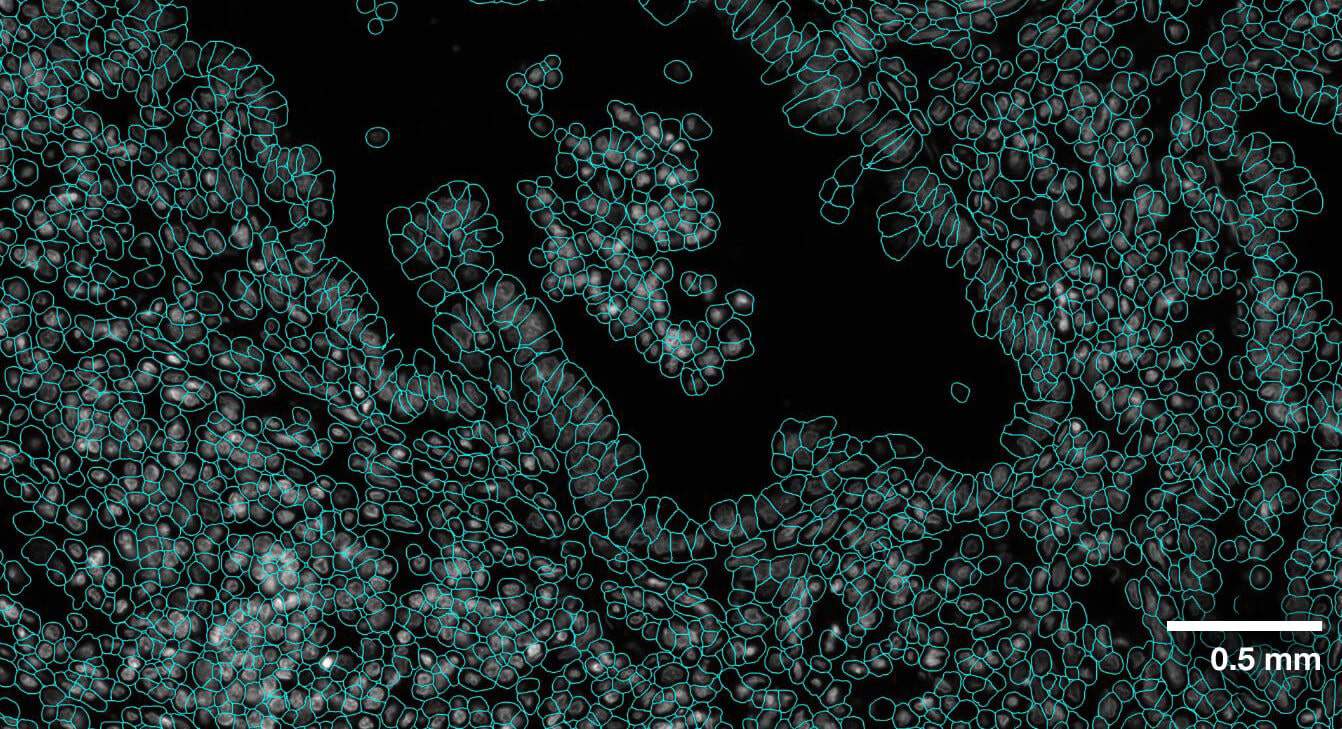
Multi-modal cell segmentation process provides accurate cell boundaries detection. CosMx cell segmentation uses cell membrane and morphology marker protein images, machine-learning augmented cell segmentation algorithm and transcript-based segmentation refinement to achieve precise single-cell segmentation in morphologically intact tissue.
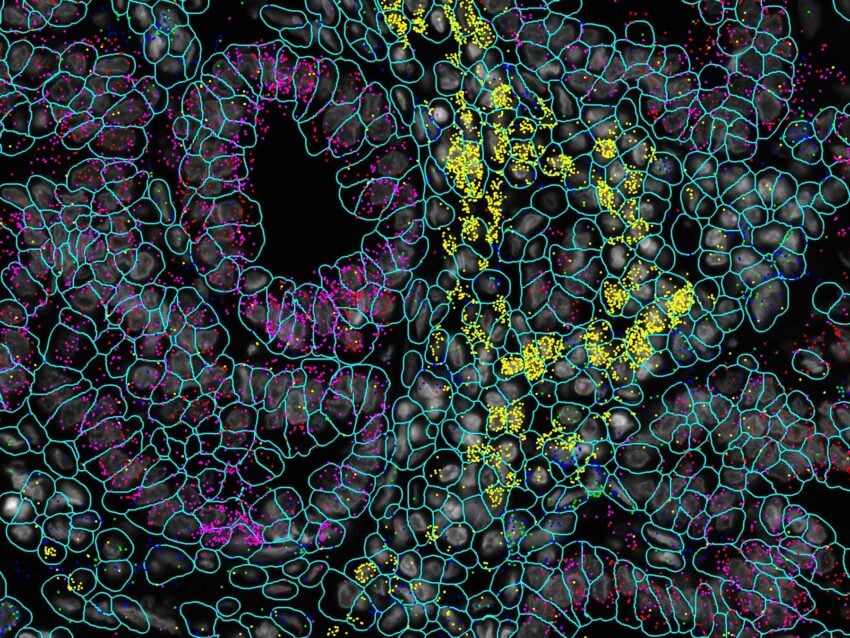
Detect up to 1000-plex RNA and 100-plex protein in intact tissue samples


In addition to RNA, detect and spatially visualize up to 100 proteins on the intact tissues. Shown here are 7 protein targets visualized on a normal colon FFPE tissue.
CosMx SMI vs. Xenium: Comparative Analysis of Breast Cancer Tissue Samples
How it works
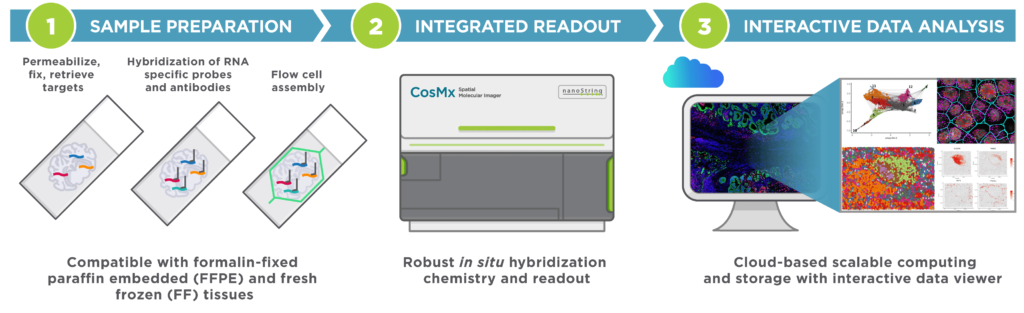
CosMx SMI is an integrated system with mature cyclic fluorescent in situ hybridization (FISH) chemistry, high-resolution imaging readout, interactive data analysis and visualization software.
Easy Sample Preparation, Compatible with Any Sample Type
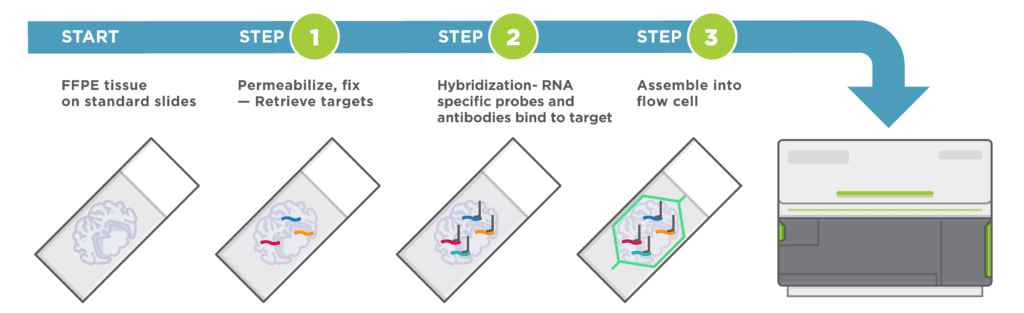
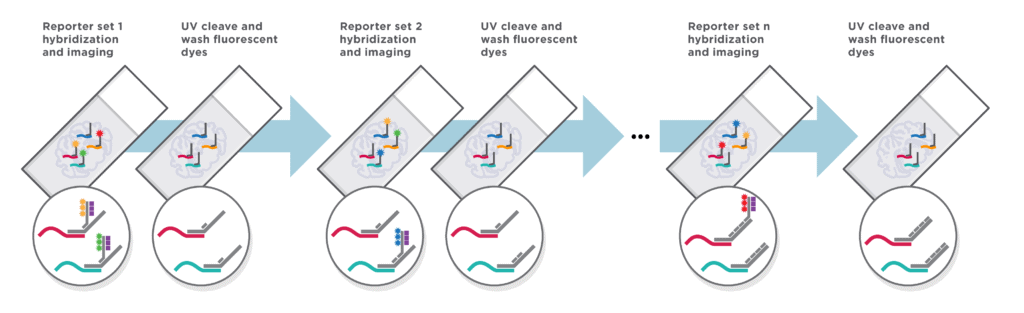
Automated Cyclic in situ Hybridization Chemistry
Applications
CosMx Spatial Molecular Imager is the most flexible and robust spatial single-cell imaging platform for:
- Cell Atlas and Characterization: Define cell types, cell states, tissue microenvironment phenotypes, and gene expression networks within spatial context.
- Cell-cell Interaction: Understand biological process controlled by ligand-receptor interactions.
- Spatial Biomarkers: Quantify change in gene expression based on treatment and identify single-cell subcellular biomarkers with spatial context.

Interested in learning more about CosMx SMI for Single-Cell Imaging?
Contact our helpful experts and we’ll be in touch soon.

Related Resources





Single-Cell Imaging FAQs
Why is a single cell important?
Cells are a fundamental unit of life. A comprehensive understanding of how cells organize themselves in different layers of information to form tissues is not yet fully achieved. Further, no matter how seemingly homogeneous a tissue might appear, it contains a diverse population of cells, all of which represent different manifestations of that tissue type. Learn more »
Why is single-cell analysis important?
Single-cell analysis encompasses the study of genomics, transcriptomics, proteomics, and metabolomics at single-cell resolution. As cells are the organism’s building blocks, they are organized in different layers of information to form tissues, and the position of each cell within a tissue has a physiological or morphological function. Learn more »
What is single-cell technique?
Single-cell techniques are advances in single-cell manipulation and amplification that have enabled the study of genomics, transcriptomics, and epigenomics at the level of a single cell. Learn more »
What is single-cell spatial transcriptomics?
Analysis of mRNA expression profile with spatial context at the level of a single cell is known as single-cell spatial transcriptomics. Each cell has a unique transcriptomic fingerprint as gene expression patterns can be heterogeneous even amongst similar cells in both standard and abnormal cell states. Learn more »
Is spatial transcriptomics single-cell resolution?
Yes. Recent advances in the field of spatial transcriptomics have made it possible to visualize RNA transcripts at the resolution of a single-cell and, in some cases, subcellular resolution. Learn more »
What is single-cell analysis used for?
Single-cell analysis can provide data on cellular phenotypes by studying the effects of genomic alterations, gene expression, and environmental influences at the level of a single cell. Learn more »