GeoMx® Mouse Whole Transcriptome Atlas
Helping Your Research
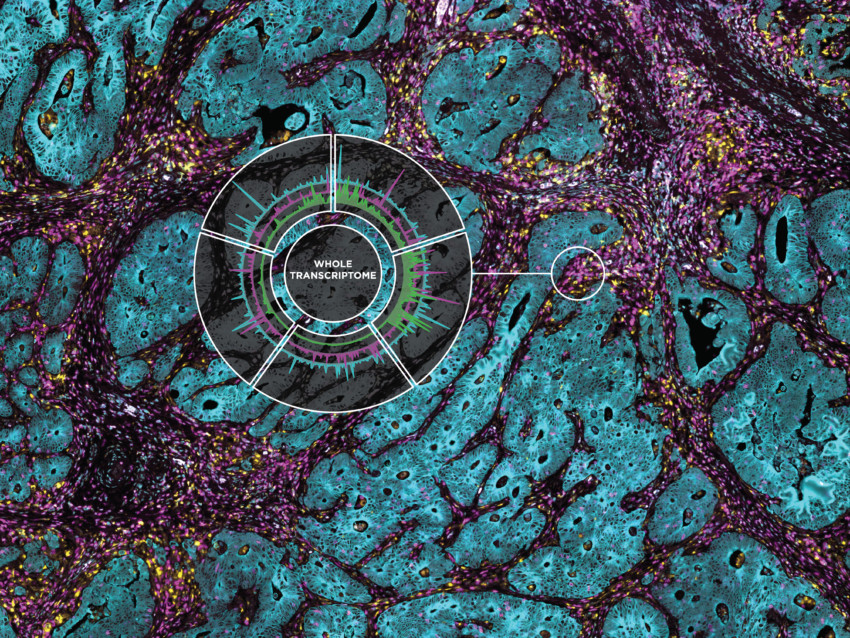
Full Transcriptome Spatial RNA Analysis
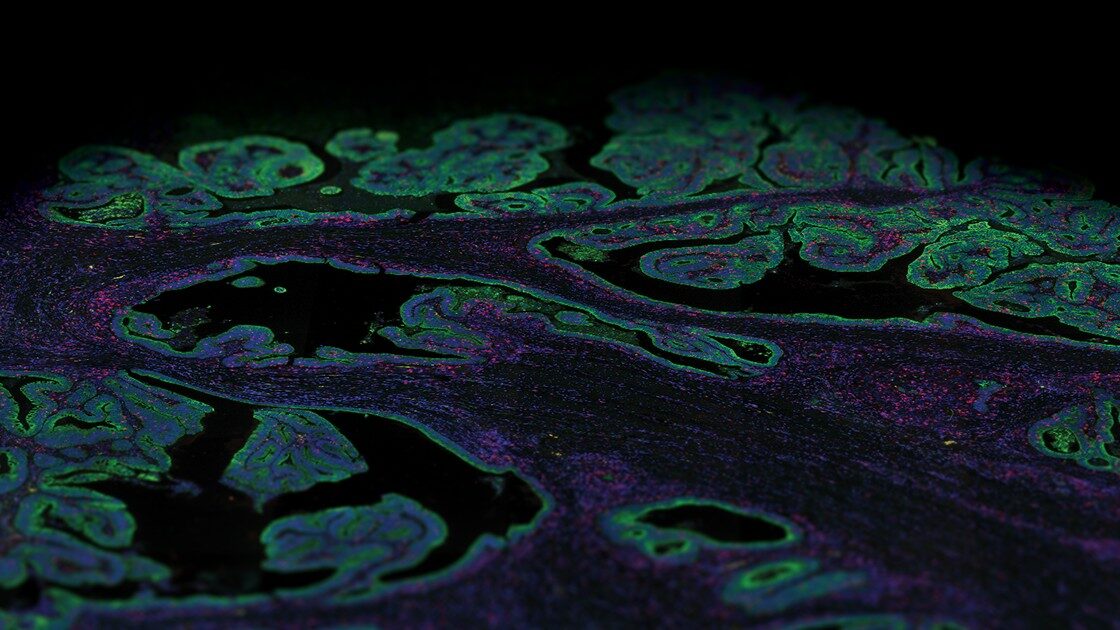
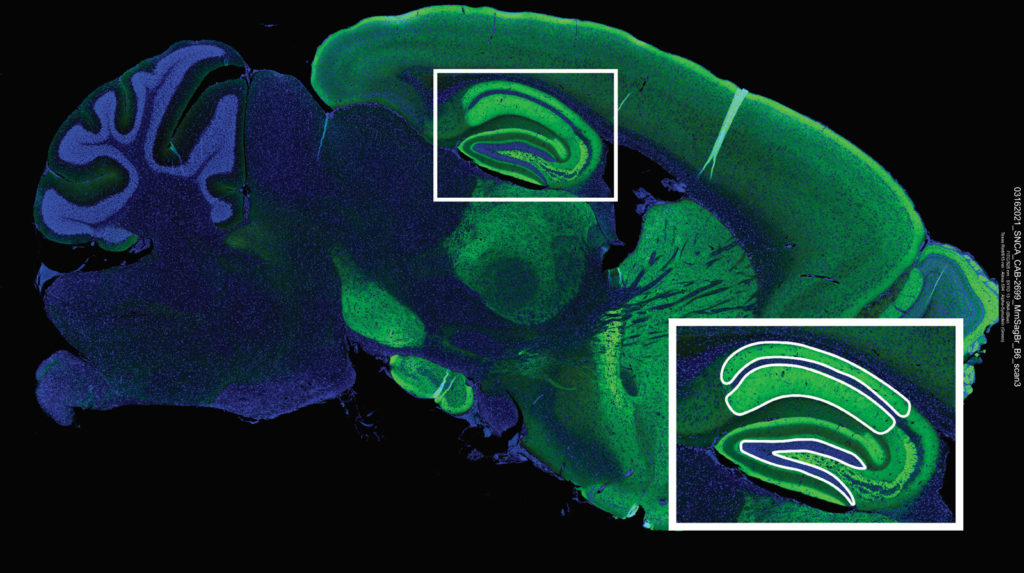
Mouse models serve a critical role in understanding developmental biology, disease onset, progression, and treatment. Tissue heterogeneity, however, confounds the results from many mouse transcriptome studies based on bulk or single cell RNA-seq data. Using the GeoMx Mouse Whole Transcriptome Assay, one can apply spatial transcriptomics to reveal the tissue architecture and underlying function in genetically modified mouse models.

Publications & Posters
Characterizing Late-Onset AD Models Using Spatial Whole Transcriptome Analysis – AGBT 2021
Smarca4-deficient lung cancers display a metastatic-like cell state and a distinct cell-of-origin – AGBT 2021
Opposing immune and genetic mechanisms shape oncogenic programs in synovial sarcoma
Synovial sarcoma (SyS) is an aggressive neoplasm driven by the SS18-SSX fusion, and is characterized by low T cell infiltration. Here, we studied the cancer-immune interplay in SyS using an integrative approach that combines single-cell RNA sequencing (scRNA-seq), spatial profiling and genetic and pharmacological perturbations.
The spatial landscape of lung pathology during COVID-19 progression
Recent studies have provided insights into the pathology and immune response to coronavirus disease 2019 (COVID-19)1–8. However, thorough interrogation of the interplay between infected cells and the immune system at sites of infection is lacking.
Temporal and spatial heterogeneity of host response to SARS-CoV-2 pulmonary infection
The relationship of SARS-CoV-2 pulmonary infection and severity of disease is not fully understood. Here we show analysis of autopsy specimens from 24 patients who succumbed to SARS-CoV-2 infection using a combination of different RNA and protein analytical platforms to characterize inter-patient and intra-patient heterogeneity of pulmonary virus infection.
Related Resources




Request a Quote
Contact our helpful experts and we’ll be in touch soon.